Protein Dynamics, Allostery, AI, and Mechanism-Guided Drug Discovery
My research group develops mathematical, computational, and AI-guided approaches to understand protein dynamics, allostery, and function, with applications in mechanism-guided drug discovery. A central goal is to predict how ligands, mutations, antibodies, and designed binders reshape allosteric communication, conformational ensembles, signaling efficacy, selectivity, and pharmacological outcomes.
A major theme of my work is to move beyond static protein structures and binding alone toward functional molecular design. Many therapeutically important proteins, including GPCRs, enzymes, signaling proteins, molecular machines, and intrinsically disordered proteins, cannot be fully understood from a single structure. Their function depends on conformational dynamics, allosteric communication, and shifts between inactive, intermediate, and active-like states. Understanding these mechanisms is essential for designing next-generation therapeutics, particularly allosteric modulators, biased ligands, and state-selective antibodies or nanobodies.
I developed Rigidity Transmission Allostery (RTA), a mathematical and computational framework for predicting long-range allosteric communication from protein structures. RTA models how local perturbations, such as ligand binding, mutation, or antibody engagement, propagate through a protein structure to alter rigidity, flexibility, and conformational degrees of freedom at distant functional sites. We integrate RTA with structural biology, NMR, HDX, functional assays, geometric Monte Carlo simulations, machine learning, and emerging AI-based design methods.
My current research sits at the interface of computational biology, structural biology, mathematical rigidity theory, artificial intelligence, and drug discovery. Our earlier work focused strongly on foundational algorithmic and mathematical method development. Increasingly, we are applying these methods to translational allosteric drug discovery, especially for GPCRs and other therapeutically important targets, in collaboration with experimental, clinical, pharma, and biotech partners.
Current Research Directions
1. Mechanism-guided allosteric drug discovery
We are developing computational and AI-guided strategies to identify functionally relevant allosteric sites, allosteric pathways, and molecular perturbations that can modulate protein function. Instead of prioritizing molecules or binders only by binding pose, affinity, or structural compatibility, our goal is to prioritize candidates by their predicted functional mechanism: how they reshape conformational ensembles, allosteric communication, efficacy, selectivity, and signaling outcomes.
This work has applications to small-molecule allosteric modulators, allosteric fragments, antibodies, single-domain antibodies/nanobodies, and designed binders. We are particularly interested in difficult-to-drug targets where active sites are conserved, poorly defined, or insufficient for achieving selectivity.
2. GPCR activation, biased signaling, and allosteric modulation
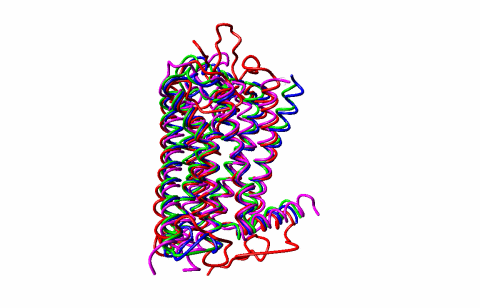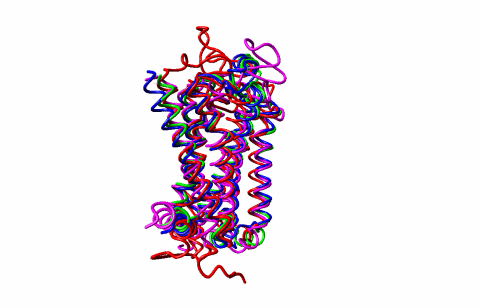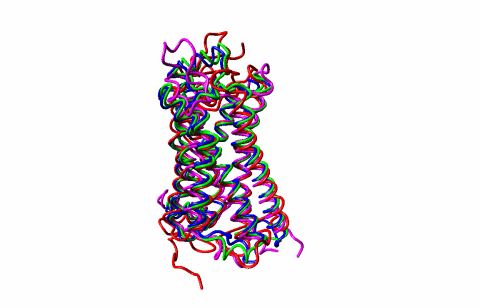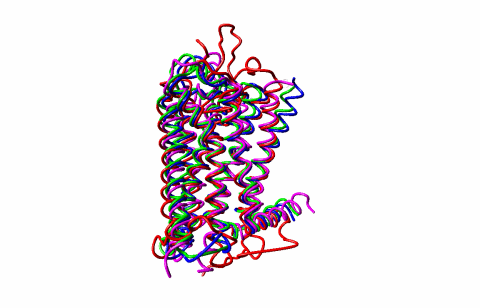
GPCRs are among the most important drug targets in medicine and are regulated by complex allosteric mechanisms. My group uses RTA, geometric simulations, NMR, structural biology, and functional assays to understand how GPCRs transition between inactive, intermediate, active, and G-protein-coupled states.
We are especially interested in how ligands, sodium ions, mutations, G proteins, antibodies, and designed binders alter GPCR allosteric networks. These insights can guide the rational design of allosteric modulators, biased ligands, and state-selective biologics with improved therapeutic precision.
3. AI-guided protein dynamics and AlphaFold/NMR/HDX structure validation
AI-based structure prediction has transformed structural biology, but static models alone do not explain protein dynamics, allostery, or function. We develop methods to evaluate, refine, and extend AlphaFold and experimentally determined structures using rigidity theory, NMR, HDX, and machine learning.
This includes structure validation and refinement of NMR ensembles, AlphaFold models, and protein complexes. Our work includes ANSURR, a rigidity-based method for NMR structure validation, and related approaches for integrating experimental constraints with computational models to improve functional structural ensembles.
4. Intrinsically disordered proteins and conformational ensembles
Intrinsically disordered proteins and regions play major roles in cancer, neurodegeneration, signaling, and phase separation, but remain difficult to characterize using conventional structure-based methods. We use geometric Monte Carlo simulations, rigidity theory, machine learning, reinforcement learning, and experimental constraints such as HDX, NMR and SAXS to model IDP conformational ensembles.
A key goal is to understand how disorder, post-translational modifications, and conformational heterogeneity influence biological function and disease mechanisms.
5. Mathematical and algorithmic foundations of allostery
My group continues to develop the mathematical and algorithmic foundations of protein dynamics and allostery. These methods draw on rigidity theory, graph theory, discrete and computational geometry, combinatorial algorithms, matroid theory, and large-scale computation.
We are developing new algorithms for proteome-scale analysis of protein flexibility, rigidity, allosteric communication, mutation effects, ensemble dynamics, and allosteric pathway prediction. These methods are designed to address major computational bottlenecks created by the rapid growth of structural biology and AI-generated protein structure data.
6. Geometry-based simulations, ensembles, and long-timescale functional dynamics
Many biologically relevant conformational changes occur on timescales that are difficult to capture using conventional molecular dynamics alone. We develop and apply geometric Monte Carlo dynamics and related accelerated sampling methods to explore functional conformational landscapes, including activation pathways, intermediate states, and ensemble shifts.
These simulations are integrated with RTA, NMR, HDX, cryo-EM, AlphaFold models, and functional experiments to connect structural ensembles with biological mechanism.
7. Broader mathematical and interdisciplinary applications
The mathematical ideas underlying rigidity, flexibility, constraint networks, and allosteric propagation also have applications beyond proteins, including mechanical linkages, robotics, metamaterials, sensor networks, and related areas of discrete applied mathematics. These foundational directions remain an important part of my broader research program and continue to inspire new methods for biological systems.
CURRENT FOCUS: mechanism-guided allosteric drug discovery and pharma/biotech collaborations | GPCR activation and biased signaling | Rigidity Transmission Allostery | AI-guided protein dynamics | NMR/HDX and experimental validation | AlphaFold and NMR structure refinement | intrinsically disordered proteins and conformational ensembles | mathematical and algorithmic methods for large-scale protein dynamics and allostery
For inro to mathematical rigidity theory see here.