Protein Dynamics, Allostery, AI, and Mechanism-Guided Drug Discovery
My research group develops mathematical, computational, and AI-guided approaches to understand protein dynamics, allostery, and function, with applications in mechanism-guided drug discovery. A major current focus is predicting how ligands, mutations, antibodies, and designed binders reshape allosteric communication, conformational ensembles, signaling efficacy, selectivity, and pharmacological outcomes, with the goal of enabling the discovery of next-generation therapeutics for currently difficult-to-drug targets.
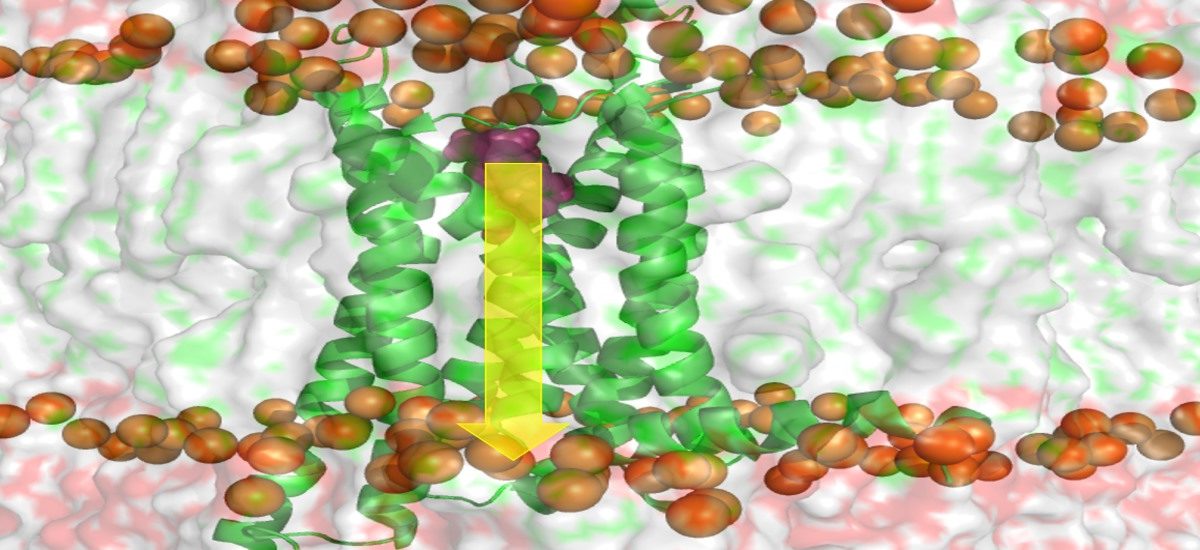
I developed Rigidity Transmission Allostery (RTA), a mathematical and computational framework for predicting long-range allosteric communication from protein structures. RTA is now being applied to translational allosteric drug discovery, particularly for GPCRs and other therapeutically important targets, by integrating structural biology, NMR, HDX, functional assays, geometric Monte Carlo simulations, machine learning, and emerging AI-based design methods to move beyond binding and static structures toward functional molecular design.
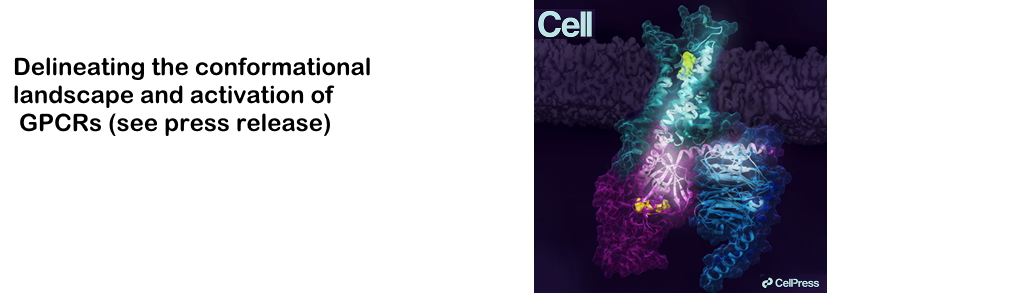
Current focus: mechanism-guided allosteric drug discovery and pharma/biotech collaborations | GPCR activation and biased signaling | AI-guided protein dynamics | NMR/HDX and experimental validation | structure validation and refinement of NMR and AlphaFold models | intrinsically disordered proteins and conformational ensembles | mathematical and algorithmic methods for large-scale protein dynamics and allostery
MY RESEARCH
MY PUBLICATIONS